-Search query
-Search result
Showing 1 - 50 of 108 items for (author: eswar & n)
EMDB-18807: 
SD1-2 fab in complex with SARS-COV-2 BA.12.1 Spike Glycoprotein.
Method: single particle / : Duyvesteyn HME, Ren J, Stuart DI
EMDB-18808: 
SD1-3 Fab in complex with SARS-CoV-2 BA.2.12.1 Spike Glycoprotein
Method: single particle / : Duyvesteyn HME, Ren J, Stuart DI

PDB-8r1c: 
SD1-2 Fab in complex with SARS-CoV-2 BA.2.12.1 Spike Glycoprotein
Method: single particle / : Duyvesteyn HME, Ren J, Stuart DI

PDB-8r1d: 
SD1-3 Fab in complex with SARS-CoV-2 BA.2.12.1 Spike Glycoprotein
Method: single particle / : Duyvesteyn HME, Ren J, Stuart DI

EMDB-41062: 
CryoEM structure of an inward-facing MelBSt at a Na(+)-bound and sugar low-affinity conformation
Method: single particle / : Guan L

PDB-8t60: 
CryoEM structure of an inward-facing MelBSt at a Na(+)-bound and sugar low-affinity conformation
Method: single particle / : Guan L

EMDB-16680: 
BA.4/5-5 FAB IN COMPLEX WITH SARS-COV-2 BA.4 SPIKE GLYCOPROTEIN
Method: single particle / : Duyvesteyn HME, Ren J, Stuart DI, Fry EE

PDB-8cin: 
BA.4/5-5 FAB IN COMPLEX WITH SARS-COV-2 BA.4 SPIKE GLYCOPROTEIN
Method: single particle / : Duyvesteyn HME, Ren J, Stuart DI, Fry EE

EMDB-26583: 
Cryo-EM structure of Antibody 12-16 in complex with prefusion SARS-CoV-2 Spike glycoprotein
Method: single particle / : Casner RG, Shapiro L

PDB-7ukl: 
Cryo-EM structure of Antibody 12-16 in complex with prefusion SARS-CoV-2 Spike glycoprotein
Method: single particle / : Casner RG, Shapiro L

EMDB-29753: 
Tomogram of an apical end of a Toxoplasma Gondii tachyzoite displaying an extruded conoid
Method: electron tomography / : Martinez M, Chang YW, Mageswaran SK

EMDB-29754: 
Cryptosporidium parvum sporozoite apical end
Method: electron tomography / : Martinez M, Chang YW, Mageswaran SK
EMDB-29755: 
Cryptosporidium parvum sporozoite basal end
Method: electron tomography / : Martinez M, Chang YW, Mageswaran SK

EMDB-29784: 
Subtomogram average of the preconoidal rings from Cryptosporidium parvum sporozoites
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29791: 
Refined subtomogram average of the top preconoidal ring from Cryptosporidium parvum sporozoites
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29801: 
Refined subtomogram average of the bottom preconoidal ring from Cryptosporidium parvum sporozoites
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW
EMDB-29808: 
Subtomogram average of the IMC surface filaments, top view, from Cryptosporidium parvum sporozoites
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29809: 
IMC surface filament, sawtooth conformation, from Cryptosporidium parvum sporozoites
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29810: 
IMC surface filament, side view, from Cryptosporidium parvum sporozoites
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29826: 
Upper preconoidal ring from Toxoplasma gondii tachyzoites
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29827: 
Lower preconoidal ring of Toxoplasma gondii tachyzoites
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29832: 
Preconoidal rings of Toxoplasma gondii tachyzoites
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29835: 
Basal IMC pore of Cryptosporidium parvum sporozoites
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29838: 
Preconoidal rings of Toxoplasma gondii tachyzoites with Formin1 conditionally depleted
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29839: 
Upper preconoidal ring of Toxoplasma gondii tachyzoites with Formin1 conditionally depleted
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-29840: 
Lower preconoidal ring from Toxoplasma gondii tachyzoites with Formin1 conditionally depleted
Method: subtomogram averaging / : Martinez M, Mageswaran SK, Chang YW

EMDB-26855: 
Arabidopsis DDM1 bound to nucleosome (H2A.W, H2B, H3.3, H4, with 147 bp DNA)
Method: single particle / : Ipsaro JJ, Adams DW, Joshua-Tor L

PDB-7ux9: 
Arabidopsis DDM1 bound to nucleosome (H2A.W, H2B, H3.3, H4, with 147 bp DNA)
Method: single particle / : Ipsaro JJ, Adams DW, Joshua-Tor L

EMDB-15882: 
Mp2Ba1 pre-pore
Method: single particle / : Marini G, Poland B, Leininger C, Lukoyanova N, Spielbauer D, Barry J, Altier D, Lum A, Scolaro E, Perez Ortega C, Yalpani N, Sandahl G, Mabry T, Klever J, Nowatzki T, Zhao JZ, Sethi A, Kassa A, Crane V, Lu A, Nelson ME, Eswar N, Topf M, Saibil HR

EMDB-15883: 
Mpf2Ba1 pore
Method: single particle / : Marini G, Poland B, Leininger C, Lukoyanova N, Spielbauer D, Barry J, Altier D, Lum A, Scolaro E, Perez Ortega C, Yalpani N, Sandahl G, Mabry T, Klever J, Nowatzki T, Zhao JZ, Sethi A, Kassa A, Crane V, Lu A, Nelson ME, Eswar N, Topf M, Saibil HR

PDB-8b6v: 
Mp2Ba1 pre-pore
Method: single particle / : Marini G, Poland B, Leininger C, Lukoyanova N, Spielbauer D, Barry J, Altier D, Lum A, Scolaro E, Perez Ortega C, Yalpani N, Sandahl G, Mabry T, Klever J, Nowatzki T, Zhao JZ, Sethi A, Kassa A, Crane V, Lu A, Nelson ME, Eswar N, Topf M, Saibil HR

PDB-8b6w: 
Mpf2Ba1 pore
Method: single particle / : Marini G, Poland B, Leininger C, Lukoyanova N, Spielbauer D, Barry J, Altier D, Lum A, Scolaro E, Perez Ortega C, Yalpani N, Sandahl G, Mabry T, Klever J, Nowatzki T, Zhao JZ, Sethi A, Kassa A, Crane V, Lu A, Nelson ME, Eswar N, Topf M, Saibil HR

EMDB-34664: 
Cryo-EM structure of the Mycobacterium tuberculosis cytochrome bcc:aa3 supercomplex and a novel inhibitor targeting subunit cytochrome cI
Method: electron tomography / : Mathiyazakan V, Gruber G

PDB-8hcr: 
Cryo-EM structure of the Mycobacterium tuberculosis cytochrome bcc:aa3 supercomplex and a novel inhibitor targeting subunit cytochrome cI
Method: electron tomography / : Mathiyazakan V, Gruber G

EMDB-15971: 
SARS-CoV-2 Delta-RBD complexed with Fabs BA.2-36, BA.2-23, EY6A and COVOX-45
Method: single particle / : Duyvesteyn HME, Ren J, Stuart DI

PDB-8bcz: 
SARS-CoV-2 Delta-RBD complexed with Fabs BA.2-36, BA.2-23, EY6A and COVOX-45
Method: single particle / : Duyvesteyn HME, Ren J, Stuart DI

EMDB-33892: 
Omicron spike trimer bound with P3E6 (Local Refinement)
Method: single particle / : Tang B, Dang S
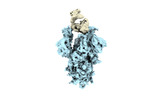
EMDB-33893: 
Omicron Spike trimer bound with up P3E6 Fab (2 RBDs up and 1 RBD down)
Method: single particle / : Tang B, Dang S
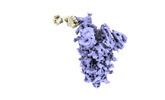
EMDB-33894: 
Omicron Spike trimer bound with side P3E6 Fab (1 RBD up and 2 RBDs down)
Method: single particle / : Tang B, Dang S

PDB-7ykj: 
Omicron RBDs bound with P3E6 Fab (one up and one down)
Method: single particle / : Tang B, Dang S

EMDB-27775: 
Cryo-EM structure of RBD-directed neutralizing antibody P2B4 in complex with prefusion SARS-CoV-2 spike glycoprotein
Method: single particle / : Reddem ER, Shapiro L

PDB-8dxs: 
Cryo-EM structure of RBD-directed neutralizing antibody P2B4 in complex with prefusion SARS-CoV-2 spike glycoprotein
Method: single particle / : Reddem ER, Shapiro L

EMDB-15073: 
Rabies virus glycoprotein in complex with Fab fragments of 17C7 and 1112-1 neutralizing antibodies
Method: single particle / : Ng WM, Fedosyuk S, English S, Augusto G, Berg A, Thorley L, Haselon AS, Segireddy RR, Bowden TA, Douglas AD

PDB-8a1e: 
Rabies virus glycoprotein in complex with Fab fragments of 17C7 and 1112-1 neutralizing antibodies
Method: single particle / : Ng WM, Fedosyuk S, English S, Augusto G, Berg A, Thorley L, Haselon AS, Segireddy RR, Bowden TA, Douglas AD

EMDB-25570: 
Cryo-EM structure of Rifamycin bound to E. coli RNAP and rrnBP1 promoter complex
Method: single particle / : Shin Y, Murakami KS

EMDB-25571: 
Cryo-EM structure of 27a bound to E. coli RNAP and rrnBP1 promoter complex
Method: single particle / : Shin Y, Murakami KS

PDB-7szj: 
Cryo-EM structure of Rifamycin bound to E. coli RNAP and rrnBP1 promoter complex
Method: single particle / : Shin Y, Murakami KS

PDB-7szk: 
Cryo-EM structure of 27a bound to E. coli RNAP and rrnBP1 promoter complex
Method: single particle / : Shin Y, Murakami KS

EMDB-24075: 
Cryo-EM structure of NTD-directed neutralizing antibody LP5 Fab in complex with SARS-CoV-2 S2P spike
Method: single particle / : Reddem ER, Casner RG, Shapiro L

PDB-7mxp: 
Cryo-EM structure of NTD-directed neutralizing antibody LP5 Fab in complex with SARS-CoV-2 S2P spike
Method: single particle / : Reddem ER, Casner RG, Shapiro L
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
